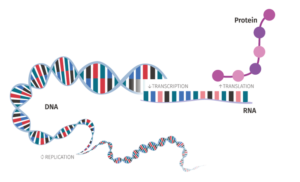
Ever wonder how gene expression is studied? What is it that makes a gene get translated into a protein a lot and others not at all? Join us and learn all about RNA and translation!
Project background
A large focus of our group is in the process of translation, how mRNA is read by ribosomes and how is the process of protein synthesis controlled. Translation occurs over the course of three distinct phases: initiation (when the ribosomes search for the translation start site and prepare for translation), elongation (the making of the protein), and termination (recognition of the stop site and disassembly of the ribosomal complex).
Translation over-activation is one of the main drivers of cancer progression. Understanding the mechanisms of translation initiation is a key step towards being able to target different types of cancers. Some treatments already target translation initiation elements to stop overactivation of translational events in cells, but more insight into the exact mechanisms is needed.
The focus for this project lies on translation initiation. Where does initiation start? How do the ribosomes scan for the initiation sites? What factors are involved in this process? How is it regulated?
Tasks
Assist and perform molecular biology laboratory experiments, including: CRISPR, PCR, gel electrophoresis, DNA & RNA extractions, qRT-PCR, Western Blotting, sequencing libraries. Depending on time commitment, this will be expanded to include: design of protocols, analysis of data, presentations in research groups.
The project involves: labwork.
Starting date/period: January 13th, 2020 – December 19th, 2020.
 Experience/skills to be acquired:
Experience/skills to be acquired:
In our lab, we use a combination of methods to study initiation of translation. On one hand, we study the effects of different known treatments and agents to understand their mechanism of action. On the other hand, we use CRISPR-Cas9 injections to mutate specific components of the translation initiation machinery in zebrafish embryos. The successful applicant will be involved in studying the effects of these treatments and genetic modifications in zebrafish. This will offer the student to learn a wide range of techniques: general zebrafish handling and development, analysis of CRISPR-guided mutant zebrafish lines, basic molecular biology techniques (gel electrophoresis, PCR, etc), qPCR experiments, RNA isolation, preparation of next-generation sequencing libraries (RNA-seq, ribosome profiling), and polysome profiling (to assess global translation events). Furthermore, the student will be made familiar with third-generation long-read sequencing on the Oxford Nanopore platform.
Involvement: Flexible, max 40 hours.
Interested by this project? Need more info? Contact Yamila Torres Cleuren (yamila.cleuren@uib.no)
Project number: 022
